Research

OctoPath research revolves around utilizing deep learning methods and medical images to develop strategies for objective diagnosis, prognosis, biomarker discovery and clinical assistance. This includes the development of AI methods for automated and objective diagnosis in histology. Integration of multimodal data, ranging from radiology scans, histology images, and multi-omics, to provide improved patient predictions, elucidate dichotomy in patient outcomes, enable in-silico treatment optimization and serve as assisting tools for clinicians. We utilize interpretability methods and quantitative analysis to explore biological relations and guide the discovery of new biomarkers. Below is overview of some ongoing and recent projects.
AI-system for Endomyocardial Biopsy Assessment
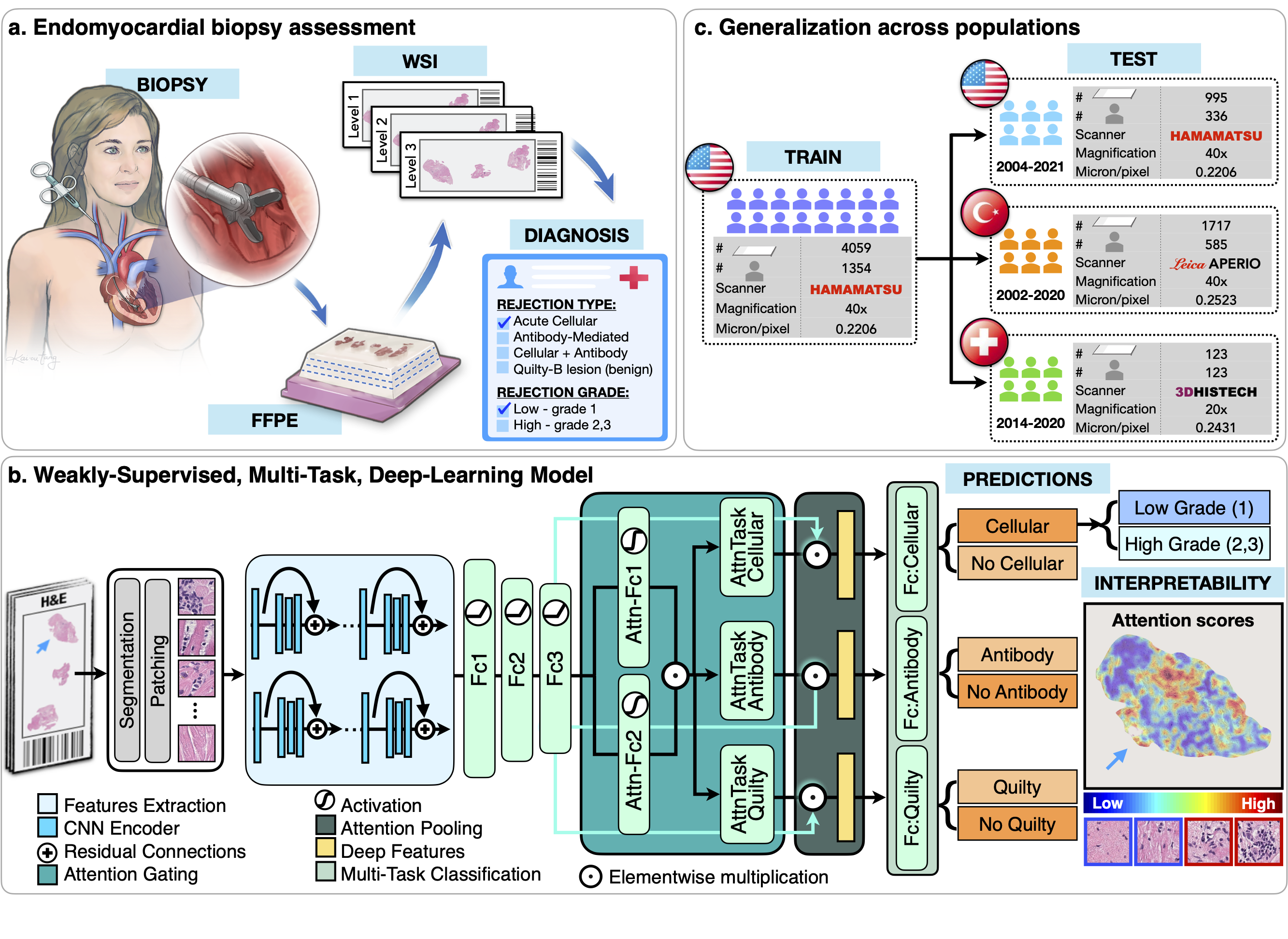
Endomyocardial biopsy (EMB) screening represents the standard of care for detecting allograft rejections after a heart transplant. Manual interpretation of EMBs is, however, affected by substantial interobserver and intraobserver variability, which often leads to inappropriate treatment with immunosuppressive drugs, unnecessary follow-up biopsies and poor transplant outcomes.
- CRANE (Cardiac Rejection Assessment Neural Estimator):
- Interpretable AI-model for simultaneous identification of heart rejection type (cellular and/or antibody-mediated), grade & benign rejection mimickers, such as Quilty-B lesions, from H&E stained whole-slide EMB images
- Multiple-instance learning is used to train model with the patient diagnosis as the only label, avoiding the need for manual feature engineering or pixel-level annotations; thus supporting large-scale deployment
- Model interpretability is obtained through attention heatmaps reflecting the relevance of each biopsy region towards the model predictions
- Model robustness is assessed on three independent cohorts from 🇺🇸, 🇹🇷 and 🇨🇭
- A reader study showed, that the model predictions are not inferior to cardiac pathologists.
- Using CRANE as assisting tool during manual biopsy reads reduces the inter-observer variability and assessment time
- journal link; demo; code
3D Printed Embedded AI-based Pathologist
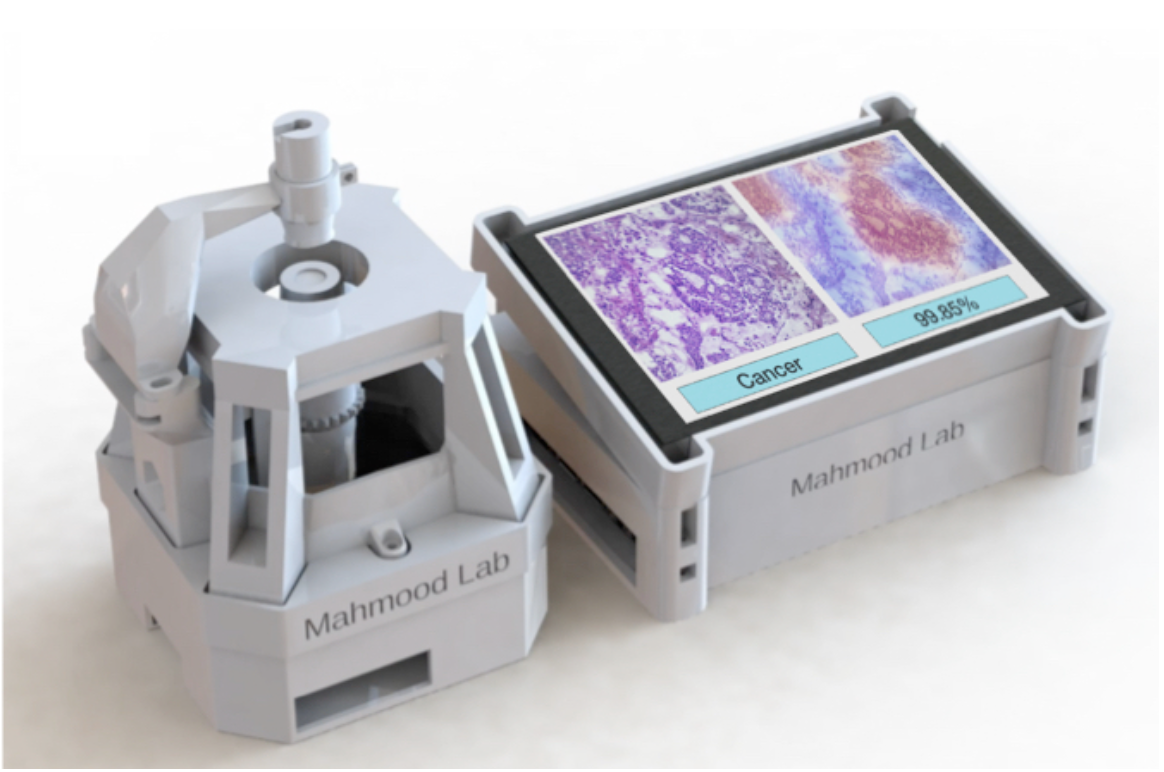
The lack of pathology diagnosis in low-to-middle income countries deprives over a billion people of necessary healthcare. AI-models could serve as possible solutions for accurate diagnosis in developing areas, however, they require expensive equipment (computing hardware, whole-slide scanner, microscopes). There is also a dire need for fast and affordable cancer diagnosis in time-critical situations such as oncologic surgery.
- We present an open source, 3D printable, cost-efficient microscope with an embedded AI model for automated diagnosis from glass pathology slides
- The system uses Raspberry Pi and camera sensor to capture high resolution (x10, x20) images of histology slides
- AI model (trained on whole-slide images) is embedded into Nvidia Jetson Nano GPU unit for automated diagnosis
- The attention heatmaps allow interpretation of the model predictions
- Tested on FFPE slides and frozen sections
- Under revision in Nature Biotechnology
Multimodal Data Fusion for Survival Prediction
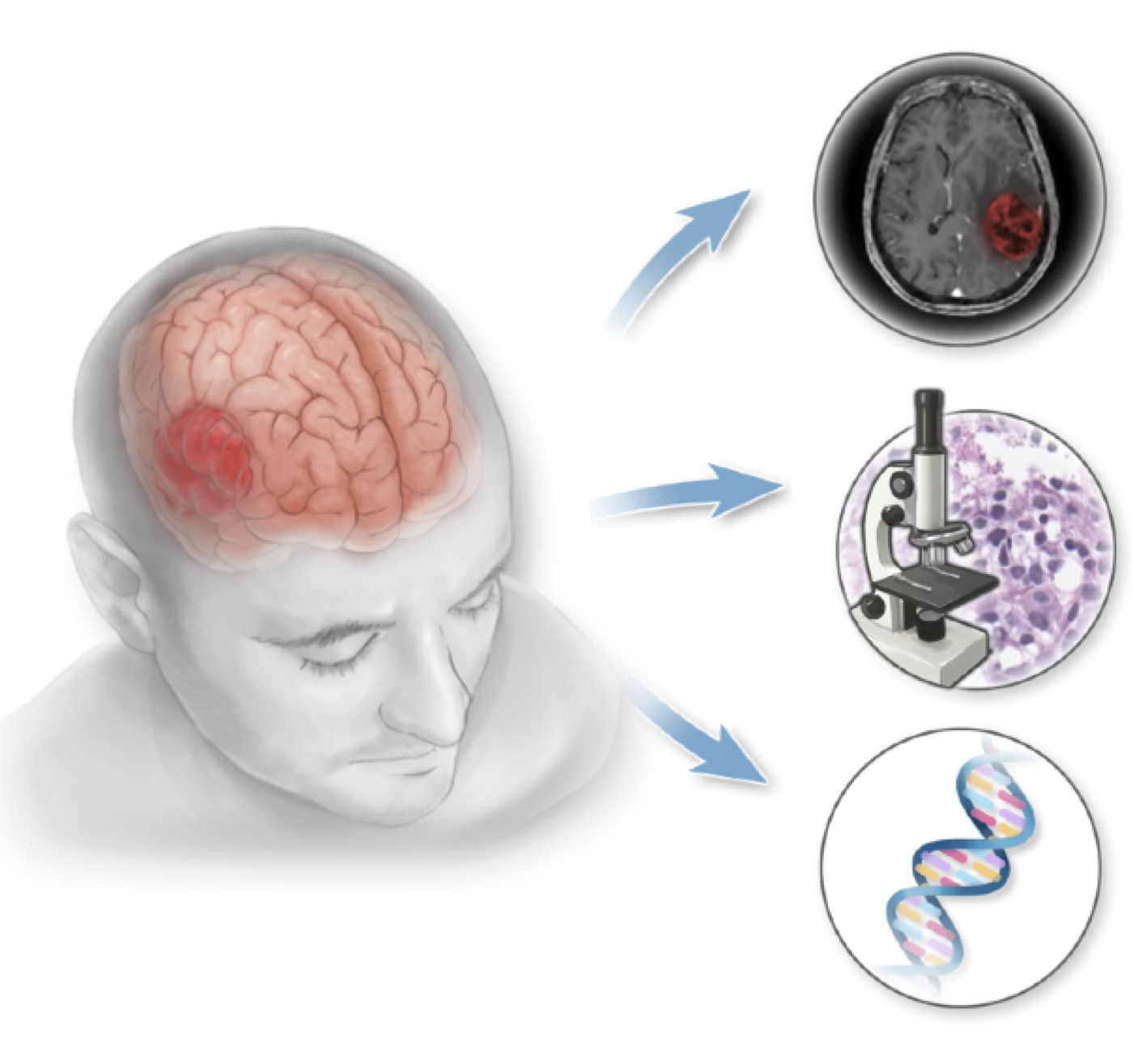
Complex processes driving glioma recurrence and treatment resistance cannot be fully understood without the integration of multiscale factors such as molecular profiles, tissue microenvironment, and macroscopic features of the tumor and the host tissue. AI holds a great potential to incorporate information from diverse data sources to provide more accurate prediction and identify novel predictive morphologies within and across modalities.
- We developed an AI-based framework for easy integration of information from multimodal data sources
- Demonstrate the feasibility of the framework by fusion of radiology, histology and genomics data for glioma survival predictions
- Weakly-supervised approach elevates the need for data annotation and supports seamless applications to other cancer models
- Transfer learning is deployed to overcome the limited size of multimodal datasets by utilizing unimodal data
- Investigation of prognostic features within and across modalities through model interpretability & shift in feature importance under unimodal vs multimodal context
- Potential for biomarker discovery and exploratory studies
- (ongoing study)
Personalized radiotherapy planning through Bayesian integration of multimodal scans
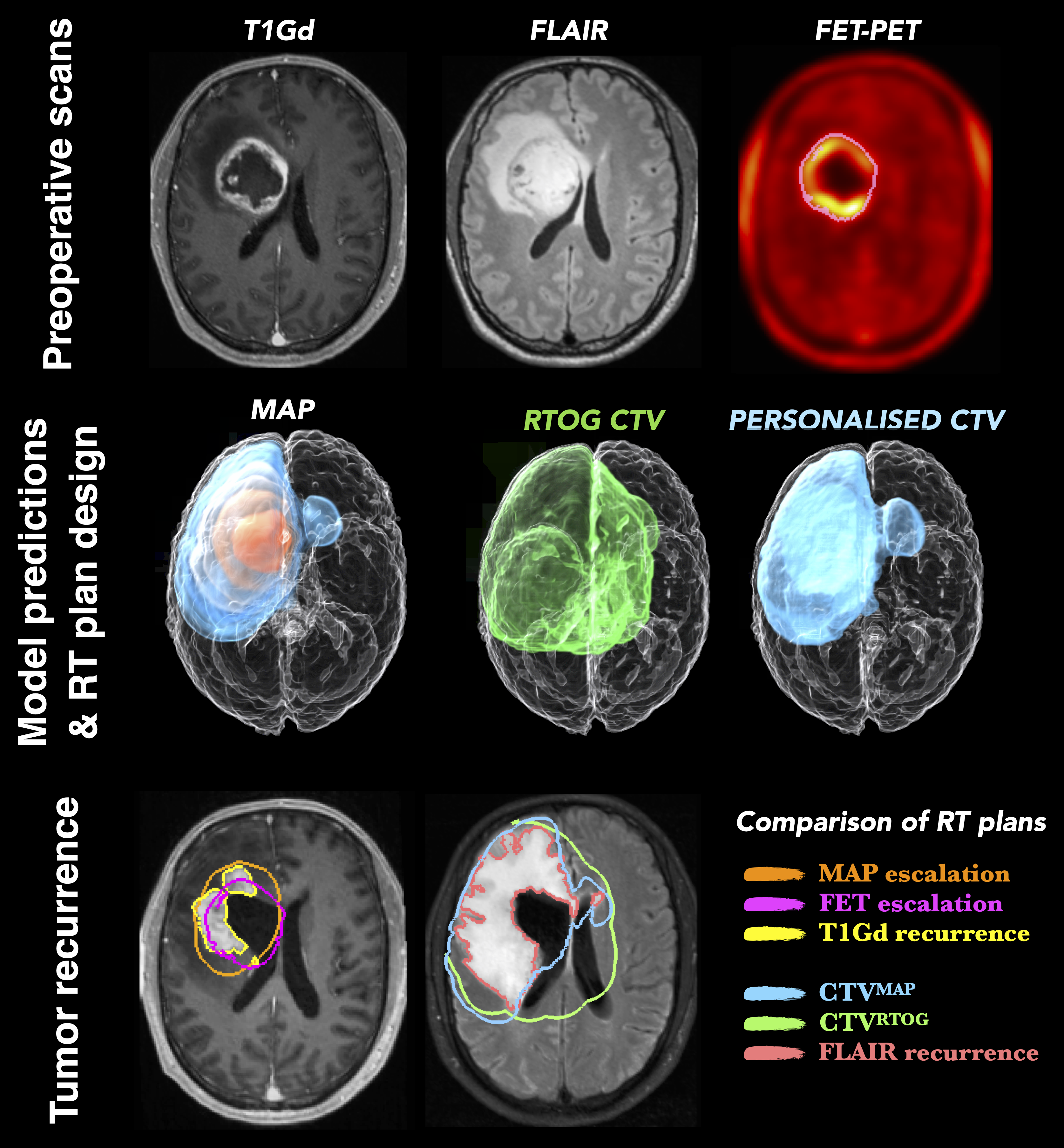
Glioblastoma (GBM) is a highly invasive brain tumor, whose cells infiltrate surrounding normal brain tissue beyond the lesion outlines visible in the current medical scans. These infiltrative cells are treated mainly by radio- therapy. Existing radiotherapy plans for brain tumors derive from population studies, assume uniform infiltration rate and scarcely account for patient-specific conditions.
- Here, we provide a Bayesian machine learning framework for the rational design of improved,
personalized radiotherapy plans using mathematical modeling and patient multimodal medical scans
- The framework integrates complementary information from structural MRI scans and FET- PET metabolic maps to infer tumor cell density in GBM patients.
- The Bayesian framework quantifies imaging and modeling uncertainties and predicts patient-specific tumor cell density with credible intervals.
- The proposed methodology relies only on data acquired at a single time point and, thus, is applicable to standard clinical settings.
- An initial clinical population study shows that the radiotherapy plans generated from the inferred tumor cell infiltration maps spare more healthy tissue thereby reducing radiation toxicity while yielding comparable accuracy with standard radiotherapy protocols
- The inferred regions of high tumor cell densities coincide with the tumor radioresistant areas, providing guidance for personalized dose- escalation
- paper; code